By Mu Huan and Eric Li
A large gene sequencing company generates 10 TB to 100 TB of offline data every day, and the big data Bioinformatics Analysis platform needs PB-level data processing capability. Biotechnology and computer technology provide two-way support behind the scenes: Sequencing applications are gradually moving from scientific research to clinical applications, computing models are evolving from offline to online, and the delivery efficiency is becoming more and more important.
James Watson and Francis Crick discovered the double helix structure of DNA in 1953. Since then, the mysteries of species evolution and heredity has been uncovered, and the human cognition of digital heredity has been opened. However, the mystery of human genes is understood bit by bit.
In 1956, a discovery of the correlation between cancer and chromosome shocked the whole cancer research community: For patients with Chronic Myeloid Leukemia (CML), chromosome 22 is significantly shorter than common chromosomes. More than two decades later, scholars discovered that the ABL gene on chromosome 9 was linked to the BCR gene on chromosome 22, and staggered translocation resulted in a BCR-Abl fusion gene. The BCR-ABL protein is always active and uncontrolled, which causes uncontrolled cell division and leads to cancer.
That is to say, as long as cells express the BCR-ABL protein, a risk of blood cancer exists. The several researchers in the United States have begun in-depth research on this, and they successfully launched a new drug for the treatment of CML. This is Glivec, also known as the "high-priced drugs" in the movie "Dying To Survive" last year.
Before the emergence of Glivec, only 30% of patients with CML survived for 5 years after diagnosis. With Glivec, this number has increased from 30% to 89%, and after 5 years, 98% of the patients have achieved complete hematological remission. For this reason, Glivec is also included in the standard list of essential drugs for WHO, and is considered to be one of the essential drugs in the medical system that are the most effective, the safest and can meet the most significant needs.
Gene sequencing is widely used in the field of hematological tumors. According to the diagnosis results of a patient, the hematological oncologist will choose corresponding examinations. For example, PCR and the real-time fluorescence probe technology is combined to detect the BCL-ABL fusion gene to diagnose CML, or SEGF (Single-end Gene Fusion) can also detect complex gene fusion types using single-end next-generation sequencing (NGS) data through the NGS technology.
On the other hand, the Non-Invasive Prenatal Testing (NIPT) for Down's (or Edwardian) syndrome, with high accuracy and low risk to the fetus, has become increasingly popular among young women in China in recent years. A gene company completes hundreds of thousands of NIPTs every year, and each NIPT involves the processing, storage and report generation for more than hundreds of MB of data. A large gene sequencing company generates 10 TB to 100 TB of offline data every day, and the big data Bioinformatics Analysis platform needs PB-level data processing capability. Biotechnology and computer technology provide two-way support behind the scenes: Sequencing applications are gradually moving from scientific research to clinical applications, computing models are evolving from offline to online, and the delivery efficiency is becoming more and more important.
In this case, a complete architecture solution is essential. Compared with Alibaba Cloud HPC, the solution should be able to split tasks as required, automatically apply for resources from the cloud, automatically scale to minimize resource holding costs, achieve a resource utilization rate of more than 90%, and automatically return computing resources after use. It maximizes resource usage efficiency, minimizes the processing cost of a single sample, and completes processing of large batches of samples in the fastest way. With the growth of gene sequencing business, it can automatically complete offline resource usage and online resource expansion. It has high speed intranet bandwidth, high throughput storage, and almost unlimited storage space.
Unlike conventional computing, Genetic computing imposes high requirements on the computing and storage capability of massive data. The computing resources on the cloud can be flexibly scheduled on a large scale mainly through the automatic scaling feature of container computing and the automatic scaling capability of Alibaba Cloud ECS. By rationally splitting genetic data, large-scale parallel computing and the TB-level sample data processing can be realized. Through the computing capability acquired on demand and the use of high-throughput object storage, the cost of holding computing resources and the processing cost of a single sample are greatly reduced.
The overall technical architecture is a hybrid cloud of cloud native containers, integrating resources on-cloud and off-cloud and managing clusters uniformly across regions. As the main player, container technology provides standardized process, acceleration, elasticity, authentication, observation, and measurement and other capabilities in data splitting, data quality control and call variation. On the other hand, high-value mining requires the use of container-based machine learning platforms and parallel frameworks to build models for large-scale linear algebra computing of genes, proteins, and medical data, thus making accurate medical capabilities a reality.
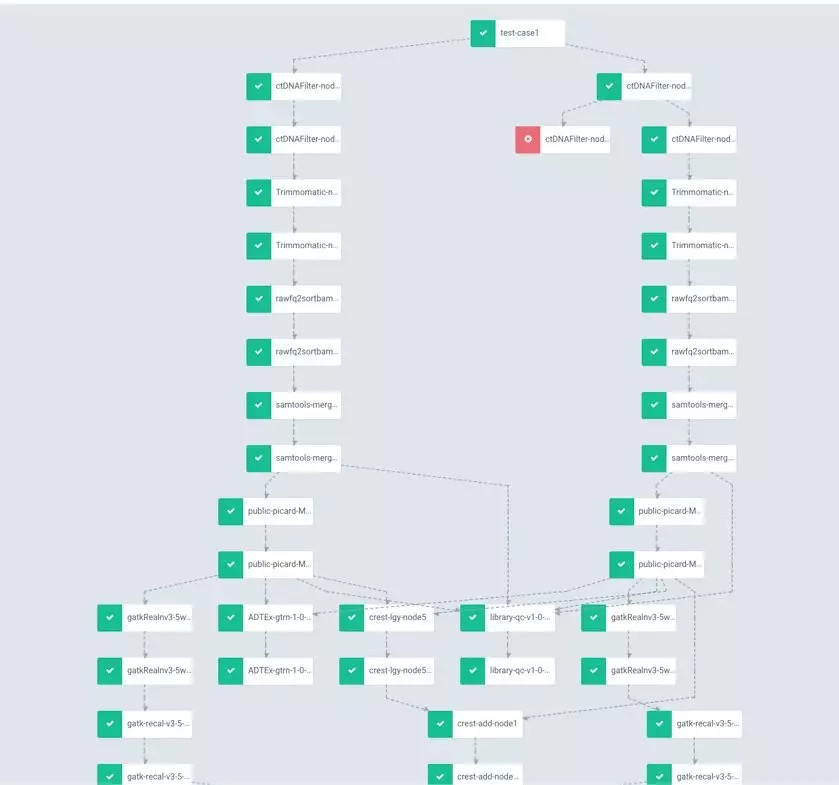
The workflow of genetic engineering is characterized by a multi-level and directed acyclic graph. DAG with a depth of 1000-5000+ for big scientific research workflow requires accurate process state monitoring and high process stability. Simple processes restart from any step. Failed steps can automatically retry and continue. Tasks, notifications, logs, audits, queries can be scheduled and operation portal CLI/UI can be integrated.
The reading of millions of small files in the data migration and data splitting phases puts pressure on the underlying file system. The processing efficiency of samples can be improved by avoiding unnecessary reading and writing of small files. Through a physical connection between the data center and Alibaba Cloud, high-throughput and low-latency data migration to the cloud, as well as cloud migration, verification, and detection combined with workflows can be implemented. The final goal is to complete the encrypted migration of dozens of TB of data in a short time, ensure the high performance and security of the data transmission client, implement concurrent transmission and resumable transmission, and maintain complete access authorization control.
A typical feature of genetic computing is that the data is computed in batches, which need to be completed sequentially according to specific steps. Abstractly, it means that a declarative workflow definition, AGS (Alibaba Cloud Genomics Service) workflow, is required.
The solution adopted is as follows:
The unified scheduling of computing resources and data aggregation on the cloud can be implemented through the hybrid cloud Alibaba Cloud Container Service for Kubernetes (ACK) cluster across IDCs and available zones on the cloud. The data migration to the cloud and subsequent data processing processes are automated and streamlined to complete the local processing, the cloud migration and the cloud-based processing for the batch offline data, and report generation within 24 hours. Computing nodes or service-free computing resources are flexibly provided on demand to form on-demand computing capability and handle sudden analysis tasks The Alibaba Cloud genetic data service team that I lead strives to build a more elastic container-based cluster with the ability to automatically scale hundreds of nodes in minutes and the Serverless capability to pull thousands of thousands of lightweight containers in minutes. It improves the utilization rate of intranet bandwidth by improving the parallelism, and ultimately improves the overall data throughput rate. It improves IO read/write speed through TCP optimization of the NAS client and server, and accelerates the reading of object storage by adding a cache layer and distributed cache for OSS.
The NovaSeq sequencer provides a low-cost ($100/WGS) and high-output (6 TB) NGS solution. The use of a large number of NovaSeq generates dozens of TB of data every day for the gene sequencing company, which requires a powerful computing capability to split and discover variations, as well as a large amount of storage to store raw data and variant data. Alibaba Cloud genetic data service continuously improves its ultimate elastic computing capabilities, massively parallel processing capabilities, and massive high-speed storage to help gene sequencing companies quickly automate the processing of dozens of TB of offline data each day, and generate high-quality variant data through the GATK standard.
With the emergence of the third generation sequencing (TGS), represented by PacBio and Nanopore, long reads from over 30 KB to several hundred KB, and a large output from 20 GB to 15 TB are generated. Long reads and large data volume request more powerful computing capability and high I/O throughput for data comparison, splitting and discovery of variation, and also bring greater challenges to optimizing genetic analysis process, splitting data, scheduling a large amount of computing resources on demand, and providing ultra-high IO throughput in the genetic computing process.
Decoding the unknown, and measuring the life. Every small step in science and technology will be a big step forward for mankind.
Eric Li, an Alibaba Cloud senior architect, a data scientist, the Top2 Winner of FDA2018 Precision Medical Competition, and a financial/biological computing industry solution expert, specializes the development of Kubernetes-based container products and the production of banking and Bioinformatics industries. Before joining Alibaba Cloud, he served as the chief architect of the Watson Data Service container platform, the architect of machine learning platform, and the gold winner of the IBM 2015 Spark global competition. He is the leader of several large-scale development projects, covering cloud computing, database performance tools, distributed architecture, biological computing, big data and machine learning.

229 posts | 34 followers
FollowAlibaba Container Service - October 15, 2024
Alibaba Developer - February 7, 2022
Alibaba Developer - September 7, 2020
Alibaba Clouder - March 6, 2020
Alibaba Clouder - April 15, 2020
Alibaba Clouder - June 5, 2020

229 posts | 34 followers
Follow Container Service for Kubernetes
Container Service for Kubernetes
Alibaba Cloud Container Service for Kubernetes is a fully managed cloud container management service that supports native Kubernetes and integrates with other Alibaba Cloud products.
Learn More ACK One
ACK One
Provides a control plane to allow users to manage Kubernetes clusters that run based on different infrastructure resources
Learn More Big Data Consulting for Data Technology Solution
Big Data Consulting for Data Technology Solution
Alibaba Cloud provides big data consulting services to help enterprises leverage advanced data technology.
Learn More Big Data Consulting Services for Retail Solution
Big Data Consulting Services for Retail Solution
Alibaba Cloud experts provide retailers with a lightweight and customized big data consulting service to help you assess your big data maturity and plan your big data journey.
Learn MoreMore Posts by Alibaba Container Service