EMR Serverless Spark notebooks support three ways to install third-party Python libraries for interactive PySpark jobs. Choose a method based on how the libraries are used and whether they need to persist across sessions.
| Method | Best for |
|---|---|
| Method 1: install with pip | Processing non-Spark variables in a notebook, such as values returned by Spark or custom variables |
| Method 2: create a runtime environment | PySpark jobs that need libraries preinstalled at every session start |
| Method 3: Add Spark configurations with a conda environment | PySpark distributed computing that requires libraries available across all Spark executors |
Prerequisites
Before you begin, ensure that you have:
A workspace. See Create a workspace
A notebook session. See Manage notebook sessions
A notebook with at least one Python cell. See Develop a notebook
Method 1: install with pip
Use this method to install libraries directly into the current session for processing non-Spark variables.
Libraries installed with pip are not persisted. Reinstall them each time you restart a notebook session.
In the E-MapReduce (EMR) console, go to EMR Serverless > Spark, click your workspace name, then click Development in the left navigation pane.
Double-click your notebook to open it.
In a Python cell, run the following command to install scikit-learn:
pip install scikit-learn
Example: classify data with scikit-learn
The following example trains a support vector machine (SVM) classifier on the Iris dataset to demonstrate scikit-learn in a notebook.
Add a Python cell, enter the following code, and click the ![]() icon to run it.
icon to run it.
# Import datasets from the scikit-learn library.
from sklearn import datasets
# Load the built-in Iris dataset.
iris = datasets.load_iris()
X = iris.data # Feature data
y = iris.target # Labels
# Split into training and test sets.
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Train an SVM classifier with a linear kernel.
from sklearn.svm import SVC
clf = SVC(kernel='linear')
clf.fit(X_train, y_train)
# Evaluate the model.
from sklearn.metrics import classification_report, accuracy_score
y_pred = clf.predict(X_test)
print(classification_report(y_test, y_pred))
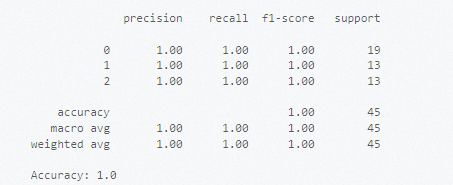
print("Accuracy:", accuracy_score(y_test, y_pred))The output looks similar to the following:
Method 2: create a runtime environment
Use this method to define a persistent custom Python environment. The libraries you specify are preinstalled each time the notebook session starts.
Step 1: create a runtime environment
In the EMR console, go to EMR Serverless > Spark, click your workspace name, then click Environments in the left navigation pane.
Click Create Environment.
On the Create Environment page, configure the Name parameter, then click Add Library in the Libraries section. For more information, see Manage runtime environments.
In the New Library dialog, set Type to PyPI, enter the library name (and optionally a version) in the PyPI Package field, then click OK. If you omit the version, the latest version is installed. Example:
scikit-learn.Click OK to create the environment. The system begins initializing the runtime environment.
Step 2: attach the runtime environment to a session
Stop the session before modifying it.
In the left navigation pane, go to O&M Center > Sessions, then click the Notebook Session tab.
Find your session and click Edit in the Actions column.
Select the runtime environment you created from the Environment drop-down list, then click Save Changes.
Click Start in the upper-right corner.
Step 3: classify data with scikit-learn
In the left navigation pane, click Development, then double-click your notebook.
Add a Python cell, enter the following code, and click the
 icon to run it.
icon to run it.
# Import datasets from the scikit-learn library.
from sklearn import datasets
# Load the built-in Iris dataset.
iris = datasets.load_iris()
X = iris.data # Feature data
y = iris.target # Labels
# Split into training and test sets.
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Train an SVM classifier with a linear kernel.
from sklearn.svm import SVC
clf = SVC(kernel='linear')
clf.fit(X_train, y_train)
# Evaluate the model.
from sklearn.metrics import classification_report, accuracy_score
y_pred = clf.predict(X_test)
print(classification_report(y_test, y_pred))
print("Accuracy:", accuracy_score(y_test, y_pred))The output looks similar to the following:

Method 3: add Spark configurations with a conda environment
Use this method when libraries must be available to Spark executors for distributed computing. You package a conda environment locally, upload it to Object Storage Service (OSS), and configure the session to use it.
Requirements: ipykernel 6.29 or later, jupyter_client 8.6 or later, Python 3.8 or later. The environment must be packaged on Linux (x86 architecture).
Step 1: create and package a conda environment
Install Miniconda:
wget https://repo.continuum.io/miniconda/Miniconda3-latest-Linux-x86_64.sh chmod +x Miniconda3-latest-Linux-x86_64.sh ./Miniconda3-latest-Linux-x86_64.sh -b source miniconda3/bin/activateCreate a conda environment with Python 3.8, install the required libraries, and package it:
# Create and activate the environment. conda create -y -n pyspark_conda_env python=3.8 conda activate pyspark_conda_env # Install third-party libraries. pip install numpy \ ipykernel~=6.29 \ jupyter_client~=8.6 \ jieba \ conda-pack # Package the environment. conda pack -f -o pyspark_conda_env.tar.gz
Step 2: upload the package to OSS
Upload pyspark_conda_env.tar.gz to an OSS bucket and copy the full OSS path. See Simple upload.
Step 3: configure and start the notebook session
Stop the session before modifying it.
In the left navigation pane, go to O&M Center > Sessions, then click the Notebook Session tab.
Find your session and click Edit in the Actions column.
On the Modify Notebook Session page, add the following to the Spark Configuration field and click Save Changes:
spark.archives oss://<yourBucket>/path/to/pyspark_conda_env.tar.gz#env spark.pyspark.python ./env/bin/pythonReplace
<yourBucket>/path/towith the OSS path you copied in the previous step.Click Start in the upper-right corner.
Step 4: process text data with jieba
jieba is a third-party Python library for Chinese text segmentation. For license information, see LICENSE.
In the left navigation pane, click Development, then double-click your notebook.
Add a Python cell, enter the following code, and click the
 icon to run it.
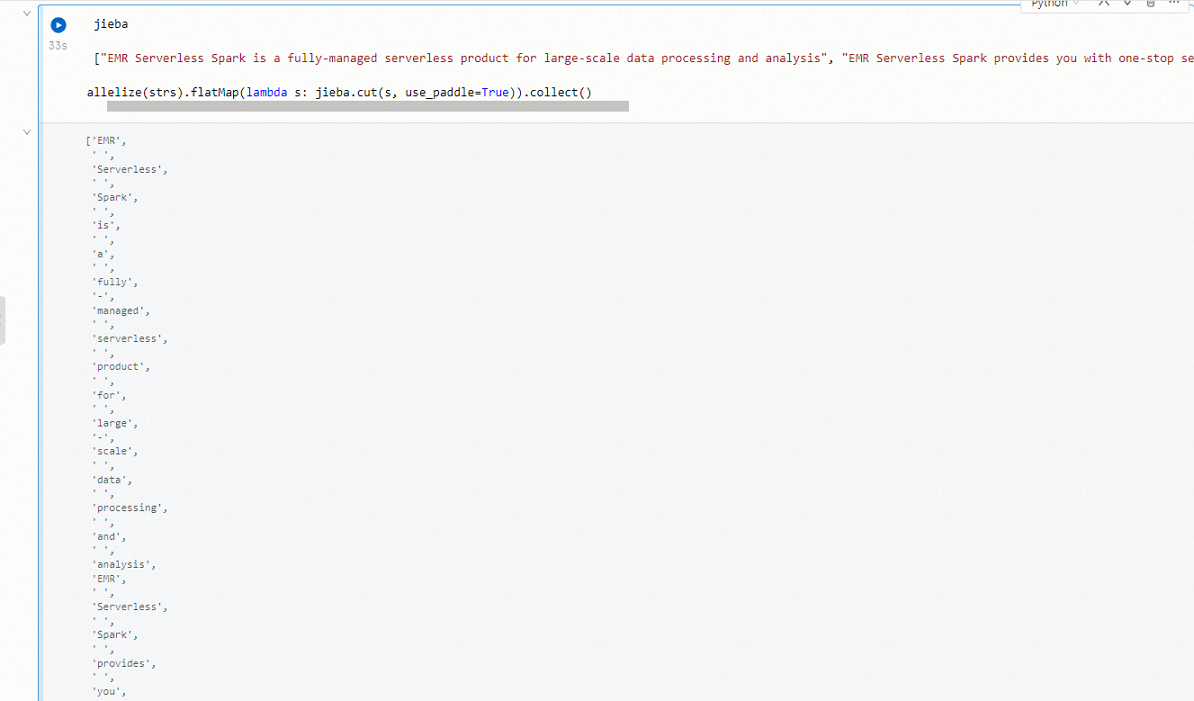
icon to run it.import jieba strs = [ "EMR Serverless Spark is a fully-managed serverless service for large-scale data processing and analysis.", "EMR Serverless Spark supports efficient end-to-end services, such as task development, debugging, scheduling, and O&M.", "EMR Serverless Spark supports resource scheduling and dynamic scaling based on job loads." ] sc.parallelize(strs).flatMap(lambda s: jieba.cut(s, use_paddle=True)).collect()The output looks similar to the following: