Use the AlphaFold3 cluster template in Elastic High Performance Computing (E-HPC) to create a cluster and deploy the Afusion web service. This document describes how to submit AlphaFold3 protein complex structure prediction jobs and view the results using Afusion.
Prerequisites
The AlphaFold3 cluster template is currently available only in the Shanghai and Beijing regions.
To enable communication between clusters, you need to create a virtual private cloud (VPC) and a vSwitch.
To enable unified storage for the cluster, you need to create a file system.
Create a cluster
Go to the Cluster List page.
Log on to the E-HPC console.
In the left part of the top navigation bar, select a region.
In the left-side navigation pane, click Cluster.
On the Cluster List page, click Cluster Template. In the dialog box that appears, select AlphaFold3.
Confirm the cluster configuration and click Create Cluster.
NoteIf an "out of stock" message appears for the selected instance type when you create the cluster, temporarily disable Queue Auto Scaling. This lets you bypass temporary resource shortages and ensures a successful cluster deployment.
To enhance system security and isolate data between different users, see User management and add a new user with
sudopermissions to avoid using therootaccount for operations.(Optional) You can configure the cluster's node auto scaling to dynamically allocate compute nodes.
Run an AF3 job
Data preparation
You can request the model parameters for this example from the official website.
Submit an AlphaFold3 job using Afusion
Log on to the Afusion page.
Go to the Cluster List page.
Log on to the E-HPC console.
In the left part of the top navigation bar, select a region.
In the left-side navigation pane, click Cluster.
On the Cluster page, find the destination cluster and click Remote Connection in the upper-right corner. The public IP address of the cluster's logon node is displayed in the Remote Connection dialog box.

Log on to the Afusion page at
http://<public_IP_of_cluster_logon_node>:8501.NoteIf the page is inaccessible, the cause is usually that port
8501is not allowed by the security group. Go to the cluster details page, switch to the Cluster Configuration tab, and click the security group ID to add a rule that allows inbound traffic on this port.
Configure the Job Settings.
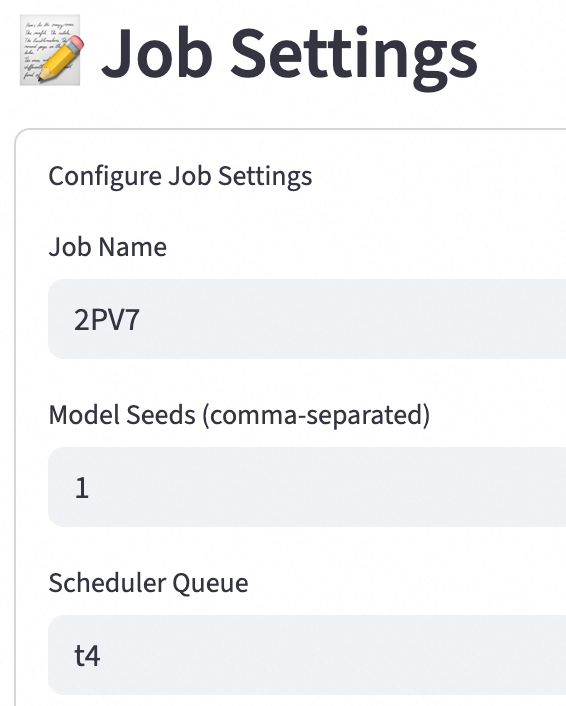
Key parameters:
Model Seeds (comma-separated): Enter one or more comma-separated numbers. These numbers are used as random seeds for the model to ensure the reproducibility of predictions. Different seeds may produce different prediction results.
Scheduler Queue: Specify the scheduling queue to which the job is submitted, such as `t4`.
Enter the biological sequence to predict. If you do not have your own sequence, you can load an open source example.
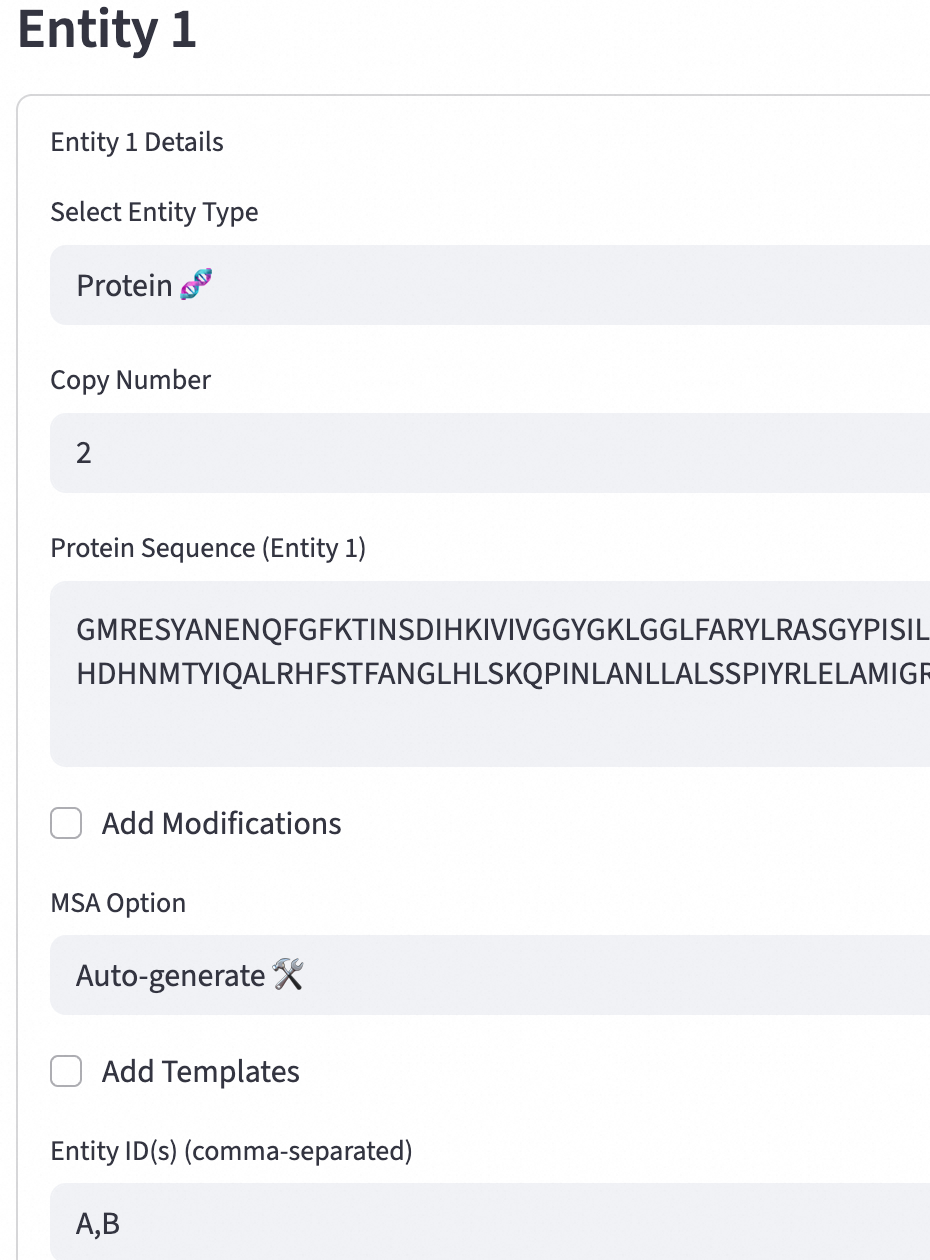
Key parameters:
Protein Sequence (Entity 1): Enter the protein sequence of the entity. This sequence is the basic data for structure prediction.
Entity ID(s) (comma-separated): A unique identifier for each entity that is used for lookup and reference in the database.
Configure the parameters required to run the AlphaFold 3 prediction and start the job.
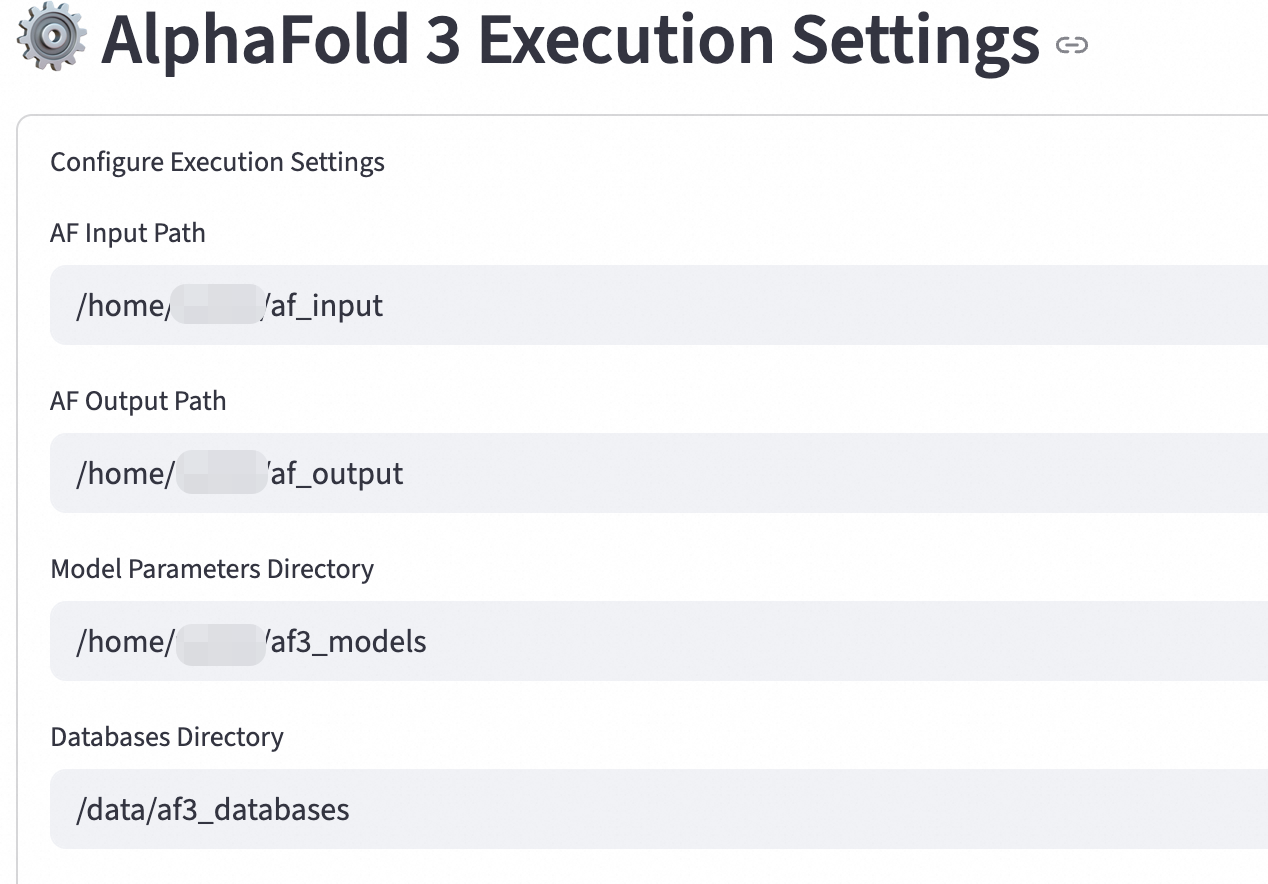

Key parameters:
AF Input Path: Specify the directory where input data is stored, such as protein sequences and multiple sequence alignments (MSAs).
Example: /home/test/af_input
AF Output Path: Specify the directory where the prediction results are saved.
Example: /home/test/af_output
Model Parameters Directory: Specify the path to the model parameter files.
Store the model parameters that you requested in the Data Preparation step in a shared directory on the cluster, such as `/opt/data` or `/home/data`.
Databases Directory: Specify the path to the database files. AlphaFold 3 uses databases such as UniRef and Pfam to perform operations, such as MSA, during prediction.
These databases are included in the E-HPC AlphaFold3 cluster template. The default path is
/data/af3_databases
After you configure the settings, click Run AlphaFold 3 Now to execute the task.
View the execution log.

View the task execution details.
In the Query AlphaFold 3 Job List section, specify the output directory for the prediction result, and click Query AlphaFold 3 Job List to query the file list.
In the Query AlphaFold 3 Job Detail section, enter the retrieved file name and click Query AlphaFold 3 Job Detail to view the details.
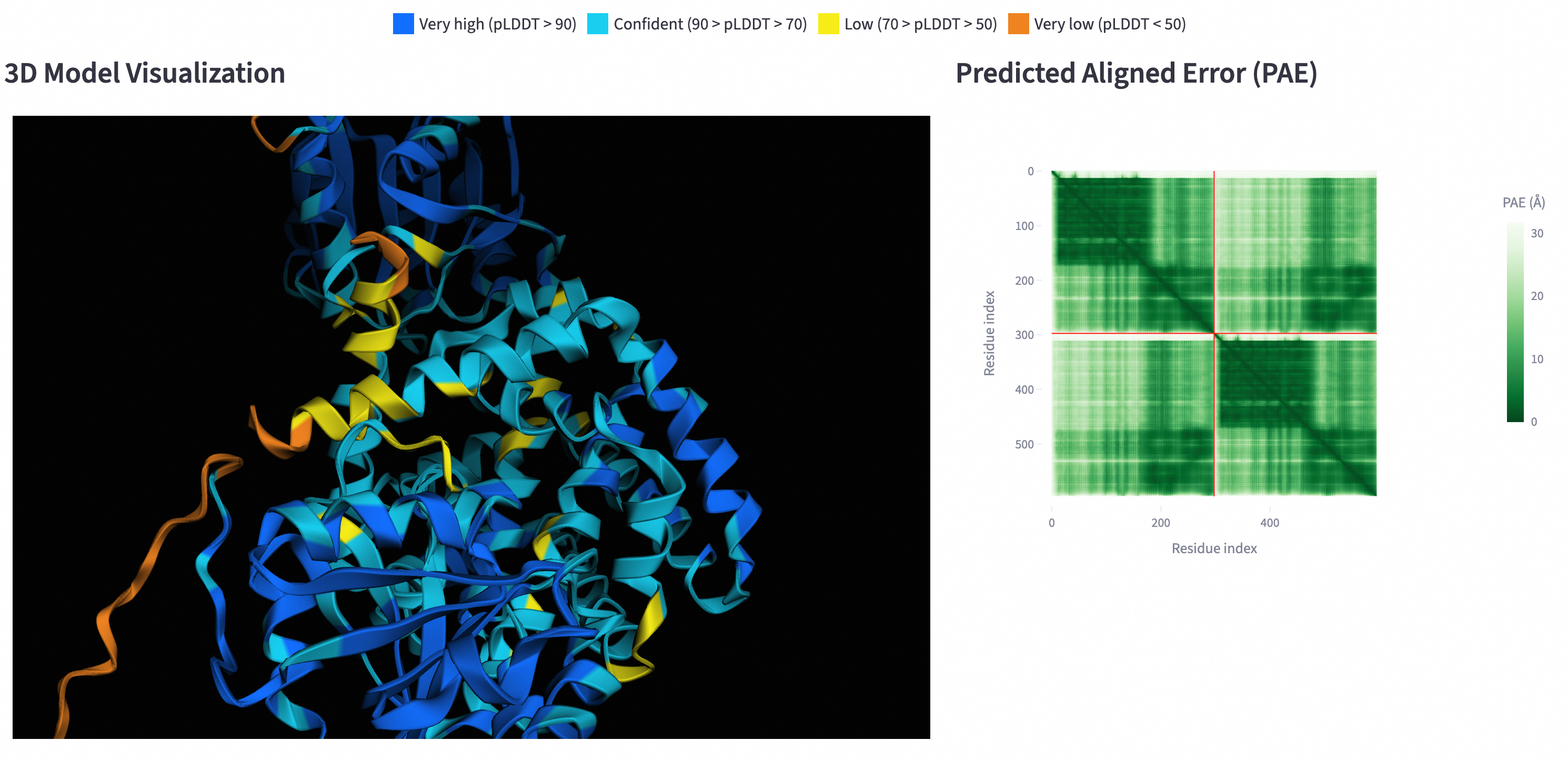
View the visualization result.
NoteIf the graph is not displayed, hardware acceleration may be disabled in your browser. For example, in Chrome, go to and enable hardware acceleration.
Download the result data. The
my_alphafold_job_model.ciffile is the main 3D protein structure file that is output by AlphaFold3. You can view it on the 3D Model Visualization page in the Afusion web interface or open it for analysis with local software such as VMD or PyMOL.
Performance reference
Test scenario: Predicting the protein folding of the 2PV7 thermophilic synthetase.
Based on the test data for this scenario, the following instance types are recommended:
For cost-effectiveness: ecs.gn7i-c8g1.2xlarge (8c30g) is recommended.
For performance: ecs.gn7i-2x.8xlarge is recommended.
This data is for reference only and does not constitute a performance commitment. Actual performance may vary depending on the specific environment, hardware, and network conditions.
Specification | GPU | Reference data storage | Inference duration (min) | Samples/hour |
ecs.gn7i-c8g1.2xlarge(8c30g) | A10 | ESSD PL0 | 31.7 | 1.89 |
ecs.gn7i-c8g1.2xlarge(8c30g) | A10 | NAS | 40.5 | 1.48 |
ecs.gn6v-c8g1.2xlarge(8c32g) | V100 | ESSD PL0 | 35 | 1.71 |
ecs.gn6i-c8g1.2xlarge(8c31g) | T4 | ESSD PL0 | 51.8 | 1.16 |
ecs.gn7i-2x.8xlarge(32c128g) | A10 | ESSD PL3 | 7.7 | 7.81 |