When cleaning pre-training corpora for large language models (LLMs) or deduplicating any large text dataset, exact matching misses near-duplicate content—paraphrased sentences, reformatted paragraphs, and partially copied text remain in the corpus and degrade model quality. Serverless Spark provides two built-in functions, minhash_lsh and build_lsh_edges, that use MinHash-Locality-Sensitive Hashing (MinHash-LSH) to identify and remove near-duplicate content at scale without the cost of pairwise comparison.
The result is approximate: the algorithm trades perfect accuracy for speed. Near-duplicate pairs with higher Jaccard similarity are more likely to be detected, but some false positives (non-duplicate pairs flagged as duplicates) and false negatives (actual duplicates that go undetected) are expected. The parameters in Performance tuning let you control this tradeoff.
How it works
The deduplication pipeline uses two stages:
-
MinHash — Converts each text document into a set of n-grams, then applies multiple hash functions to produce a compact signature vector that preserves pairwise Jaccard similarity.
-
Locality-Sensitive Hashing (LSH) — Divides the signature vector into bands and hashes each band independently. Documents that land in the same hash bucket for any band are candidate near-duplicates.
Both functions run on the Fusion engine with vectorized execution, which eliminates row-to-column conversion overhead and improves throughput.
Pipeline overview:
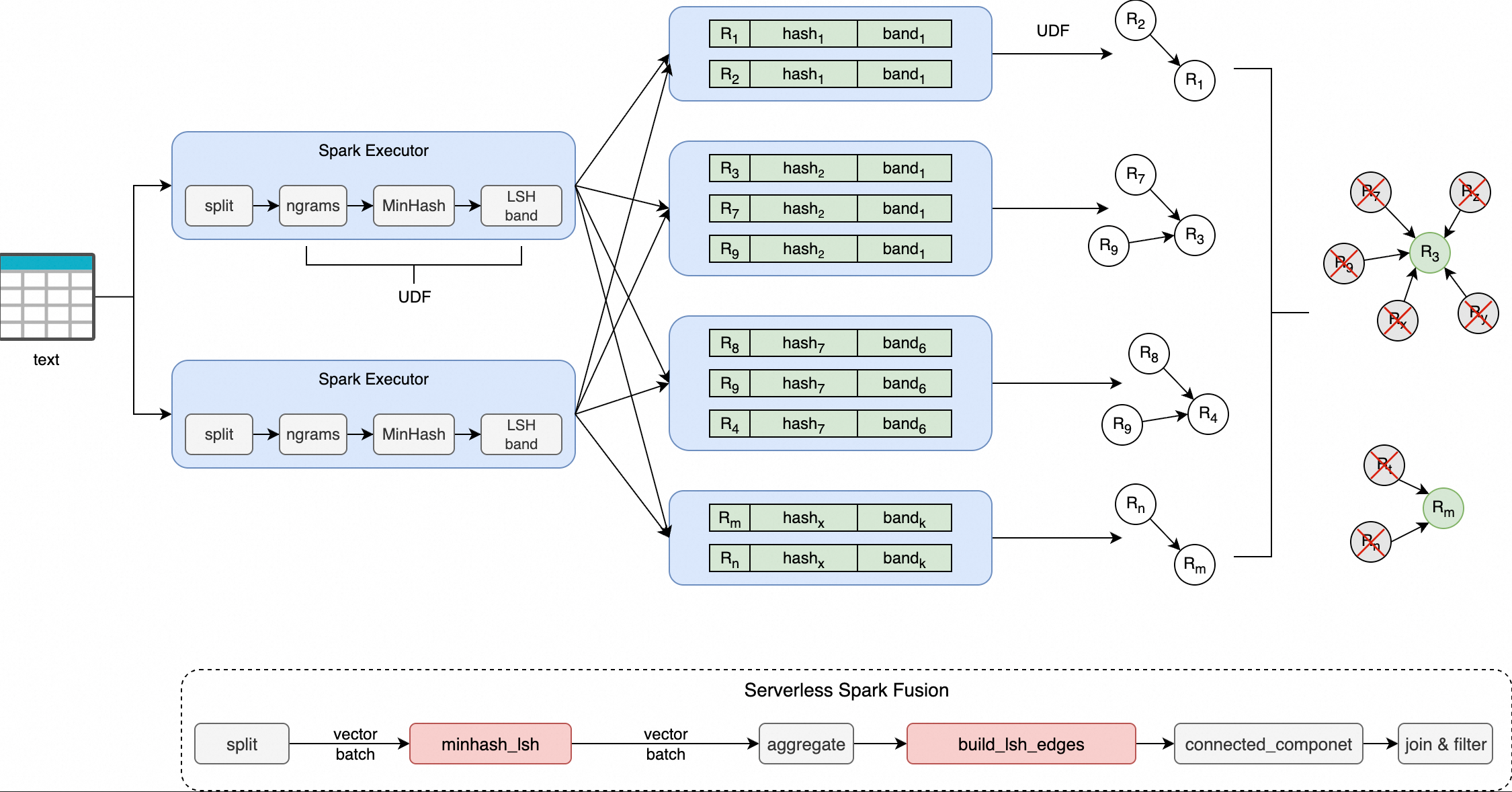
-
minhash_lsh: Takes tokenized text and produces an array of band hash values. -
build_lsh_edges: Takes the document IDs in each hash bucket and produces a graph edge list using a "minimum node connection" policy. -
After building the edge list, run graph connected components analysis to cluster near-duplicates, then keep one representative document per cluster.
Built-in functions
minhash_lsh
Tokenizes the input text, generates a MinHash signature, divides the signature into bands, and returns a hexadecimal hash value for each band.
Syntax
minhash_lsh(
tokens: ARRAY<STRING>,
perms_a: ARRAY<BIGINT>,
perms_b: ARRAY<BIGINT>,
hash_ranges: ARRAY<INT>,
ngram_size: INT,
min_length: INT
)Parameters
| Parameter | Type | Required | Description |
|---|---|---|---|
tokens |
ARRAY<STRING> |
Yes | Array of tokens after tokenization. For example, generated by split(lower(text), '\\s+'). |
perms_a |
ARRAY<BIGINT> |
Yes | Multiplier parameter *a*ᵢ for the MinHash hash function group, satisfying (*a*ᵢ · *x* + *b*ᵢ) mod *p*. |
perms_b |
ARRAY<BIGINT> |
Yes | Addend parameter *b*ᵢ for the MinHash hash function group. |
hash_ranges |
ARRAY<INT> |
Yes | Band boundaries in the format [0, R, 2R, ..., B*R], where B is the number of bands and R is the number of rows per band. |
ngram_size |
INT |
Yes | Size of each n-gram. For long texts, 5–9 is recommended. |
min_length |
INT |
Yes | Minimum number of tokens required. Documents with fewer tokens are skipped. |
Return value
-
Type:
ARRAY<STRING> -
An array of B hexadecimal strings, one per band.
-
Example:
["a1b2c3", "d4e5f6", ...]
build_lsh_edges
Takes a list of document IDs that fall into the same LSH hash bucket and generates a graph edge list using a "minimum node connection" policy: the node with the smallest ID connects to all other nodes in the bucket. This edge list is used for graph connected components analysis to cluster near-duplicate documents.
Syntax
build_lsh_edges(doc_ids: ARRAY<BIGINT>)Parameters
| Parameter | Type | Required | Description |
|---|---|---|---|
doc_ids |
ARRAY<BIGINT> |
Yes | List of document IDs in the same LSH bucket. |
Return value
-
Type:
ARRAY<STRUCT<src: LONG, dst: LONG>> -
A list of edges where the node with the smallest ID is the source and all other nodes are destinations.
-
Example: For IDs
[1003, 1001, 1005], the output is edges(1001, 1003)and(1001, 1005).
Supported versions
| Engine series | Minimum version |
|---|---|
| esr-4.x | esr-4.1.1 |
| esr-3.x | esr-3.1.1 |
| esr-2.x | esr-2.5.1 |
Example: Deduplicate the fineweb-edu dataset
This example deduplicates the sample/10BT subset of the fineweb-edu open-source dataset.
Prerequisites
Before you begin, ensure that you have:
-
An OSS bucket with write access
-
An EMR Serverless Spark workspace
-
A resource queue configured in the workspace. For setup instructions, see Manage resource queues.
Step 1: Prepare the data
-
Download the dataset. The full sample/10BT subset is 28.5 GB. This example uses a 2.15 GB portion.
-
Upload the dataset to OSS. Upload the Parquet files to a path in your OSS bucket using the console or CLI. For instructions, see Simple upload.
-
Verify the data is readable by creating a temporary view in the Spark SQL environment.
CREATE OR REPLACE TEMPORARY VIEW temp_source USING parquet OPTIONS (path 'oss://<bucket>/fineweb-edu/sample/10BT/') ;The dataset contains 727,000 records.
Step 2: Write the deduplication script
Save the following code as MinHash.py. The script uses minhash_lsh to generate band hashes, build_lsh_edges to build a graph edge list, and GraphFrames connected components to cluster and remove duplicates.
import re
from typing import List
from typing import Tuple
import pyspark
import numpy as np
import numpy.typing as npt
from graphframes import GraphFrame
from pyspark.sql import SparkSession
from pyspark.sql import functions as sf
from scipy.integrate import quad as integrate
RNG = np.random.RandomState(42)
SPLIT_PATTERN = re.compile(r"[\s\xA0]+") # Text delimiter pattern
DTYPE = np.uint32
MAX_HASH = 4_294_967_295 # Maximum 32-bit unsigned integer
MOD_PRIME = 4_294_967_291 # Largest 32-bit prime number
# Replace the placeholders below with your actual OSS paths
input_files = "oss://<bucket>/fineweb-edu/sample/10BT/*.parquet" # Input data path (Parquet format)
output_path = "oss://<bucket>/fineweb-edu/output/10BT" # Output path for deduplicated results
checkpoint_dir = "oss://<bucket>/fineweb-edu/checkpoints" # Intermediate state storage for graph computation
threshold = 0.8 # Similarity threshold. 0.7–0.9 recommended.
num_perm = 256 # MinHash signature length. Affects accuracy and performance.
ngram_size = 5 # n-gram size. 5–9 recommended for long text.
min_length = ngram_size # Minimum token count. Documents shorter than this are skipped.
text_column = "text" # Name of the text column to deduplicate
index_column = "__id__"
# Fix default parallelism to ensure stable connected components results
conf = (
pyspark.SparkConf()
.set("spark.default.parallelism", "200")
)
spark = SparkSession.Builder() \
.appName("MinHashLSH") \
.config(conf=conf) \
.enableHiveSupport() \
.getOrCreate()
spark.sparkContext.setCheckpointDir(checkpoint_dir)
# Compute optimal band parameters (B bands, R rows per band) from threshold and num_perm
def optimal_param(
threshold: float,
num_perm: int,
false_positive_weight: float = 0.5,
false_negative_weight: float = 0.5
):
def false_positive_area(threshold: float, b: int, r: int):
a, _ = integrate(lambda s: 1 - (1 - s**r)**b, 0.0, threshold)
return a
def false_negative_area(threshold: float, b: int, r: int):
a, _ = integrate(lambda s: 1 - (1 - (1 - s**r)**b), threshold, 1.0)
return a
min_error = float("inf")
opt = (0, 0)
for b in range(1, num_perm + 1):
max_r = int(num_perm / b)
for r in range(1, max_r + 1):
fp = false_positive_area(threshold, b, r)
fn = false_negative_area(threshold, b, r)
error = fp * false_positive_weight + fn * false_negative_weight
if error < min_error:
min_error = error
opt = (b, r)
return opt
# B (bands) and R (rows per band), where B x R = num_perm
# You can also set B and R manually instead of computing them from threshold.
B, R = optimal_param(threshold, num_perm)
HASH_RANGES_SLICE: List[int] = [i * R for i in range(B + 1)]
PERMUTATIONS: Tuple[npt.NDArray[DTYPE], npt.NDArray[DTYPE]] = (
RNG.randint(1, MOD_PRIME, size=(num_perm,), dtype=DTYPE),
RNG.randint(0, MOD_PRIME, size=(num_perm,), dtype=DTYPE),
)
# Assign a numeric index to each document
# If your data already has a primary key column, use it directly:
# .withColumn(index_column, sf.col("primary_key_col"))
df = spark.read.parquet(input_files) \
.withColumn(index_column, sf.monotonically_increasing_id())
a, b = PERMUTATIONS
# Step 1: Generate band hashes with minhash_lsh, then expand into (band_idx, band_hash) rows
hash_df = df \
.select(index_column,
sf.split(
sf.lower(text_column),
pattern=SPLIT_PATTERN.pattern).alias("tokens")) \
.select(index_column,
sf.minhash_lsh(
"tokens",
a.tolist(),
b.tolist(),
HASH_RANGES_SLICE,
ngram_size,
min_length).alias("hashes")) \
.select(index_column, sf.posexplode("hashes").alias("band_idx", "band_hash"))
# Step 2: For each LSH bucket (same band_idx + band_hash), generate graph edges
# using build_lsh_edges: the node with the smallest ID connects to all others in the bucket
edges_df = hash_df.groupBy("band_idx", "band_hash") \
.agg(sf.count(index_column).alias("cnt"), sf.collect_list(index_column).alias("doc_ids")) \
.filter(sf.col("cnt") > 1) \
.select(sf.build_lsh_edges("doc_ids").alias("edges")) \
.select(sf.explode("edges").alias("edge")) \
.selectExpr("edge.src as src", "edge.dst as dst") \
.persist(pyspark.StorageLevel.DISK_ONLY)
# Step 3: Build the vertex set from all edge endpoints
vertices_df = edges_df.select(sf.col("src").alias("id")) \
.union(edges_df.select(sf.col("dst").alias("id"))) \
.distinct() \
.repartition(4096) \
.persist(pyspark.StorageLevel.MEMORY_AND_DISK_DESER)
# Step 4: Run connected components to cluster near-duplicate documents
assignment = GraphFrame(vertices_df, edges_df).connectedComponents()
# Step 5: Keep only the representative document (smallest ID) from each cluster
df = df.join(assignment.select(sf.col("id").alias(index_column), sf.col("component").alias("__component__")),
on=index_column, how="left") \
.filter(sf.col("__component__").isNull() | (sf.col("__component__") == sf.col(index_column))) \
.drop("__component__")
# Write deduplicated output
df.write.parquet(output_path, mode="overwrite", compression="snappy")
# Release cached data
edges_df.unpersist()
vertices_df.unpersist()Script parameters
| Parameter | Example value | Description |
|---|---|---|
input_files |
"oss://<bucket>/fineweb-edu/sample/10BT/*.parquet" |
Input data path (Parquet format) |
output_path |
"oss://<bucket>/fineweb-edu/output/10BT" |
Output path for deduplicated results |
checkpoint_dir |
"oss://<bucket>/fineweb-edu/checkpoints" |
Storage path for intermediate graph computation state |
threshold |
0.8 |
Similarity threshold (0.7–0.9 recommended) |
num_perm |
256 |
MinHash signature length. Higher values improve accuracy at the cost of performance. |
ngram_size |
5 |
n-gram size (5–9 recommended for long text) |
min_length |
5 |
Minimum token count. Documents with fewer tokens are skipped. |
text_column |
"text" |
Name of the text column to deduplicate |
Step 3: Upload the script
-
Go to the resource upload page.
-
Log on to the E-MapReduce console.
-
In the left navigation pane, choose EMR Serverless > Spark.
-
On the Spark page, click the name of your workspace.
-
In the left navigation pane, click Artifacts.
-
-
On the Artifacts page, click Upload File.
-
In the Upload File dialog box, click the upload area or drag
MinHash.pyinto it.
Step 4: Create and run a batch job
-
On the EMR Serverless Spark page, click Development in the left navigation pane.
-
On the Development tab, click the
 icon.
icon. -
Enter a name, set Type to Application(Batch) > PySpark, and click OK.
-
In the upper-right corner, select a resource queue.
-
Configure the following settings, leave all other parameters at their defaults, and click Run.
Parameter Setting Main Python Resource Select MinHash.pyfrom the Artifacts page.Engine Version esr-2.8.0,esr-3.4.0,esr-4.4.0, or later.Resource Configuration 4 CPU cores to 16 GB memory. For example: spark.executor.cores=4, spark.executor.memory=14GB, spark.executor.memoryOverhead=2GB.Spark Configuration Required: spark.rdd.ensureConfigConsistency true— must be set totrue.<br><br>Execution efficiency:<br>-spark.sql.shuffle.partitions 1000— for datasets under 1 TB. Add 1,000 per additional 1 TB.<br>-spark.sql.files.maxPartitionBytes 256MB— controls partition size during the read phase.128MBor256MBis typical. -
After the job starts, go to Execution Records at the bottom of the page and click Details to monitor progress.
Step 5: Verify the results
Create a temporary view to load the deduplicated output.
CREATE OR REPLACE TEMPORARY VIEW
temp_target USING parquet OPTIONS (path 'oss://<bucket>/fineweb-edu/output/10BT')
;The output contains 724,809 records, down from the original 727,000. This means 2,191 near-duplicate documents were removed.
Performance tuning
Tune the following parameters to balance precision, recall, and execution efficiency for your data scale and requirements.
Algorithm parameters
| Parameter | Recommended value | Description |
|---|---|---|
num_perm |
128 or 256 | Higher values increase accuracy but also increase computation time. 256 is sufficient for most cases. |
ngram_size |
5–9 for long text; 2–3 for short sentences | Controls semantic granularity. Values that are too large reduce sensitivity. |
threshold |
0.7–0.9 | Values above 0.9 may produce insufficient recall. |
LSH band configuration: B and R
The MinHash signature is divided into B bands, each containing R rows, where B x R = num_perm. This determines the shape of the LSH probability curve P(s) = 1 - (1 - s^R)^B, which controls how sharply the detection probability rises near the similarity threshold.
Why B and R matter:
-
More bands (larger B, smaller R): each band covers fewer rows, so two documents need to match in only one of many bands to be flagged as candidates. More bands increase the probability of any single match, which raises recall (fewer missed duplicates) but also increases false positives.
-
Fewer bands (smaller B, larger R): each band must match fully across more rows, requiring stronger similarity evidence before flagging. This raises precision (fewer over-deletions) but may miss borderline duplicates.
Recommended configurations for `num_perm=256`
| Goal | B (number of bands) | R (rows per band) |
|---|---|---|
| High recall (minimize missed duplicates) | 32 | 8 |
| High precision (minimize over-deletion) | 8 | 32 |
| Balanced (recommended starting point) | 16 | 16 |
Spark execution optimization
| Configuration | Recommended setting | Description |
|---|---|---|
spark.sql.shuffle.partitions |
1,000 for datasets under 1 TB; add 1,000 per additional 1 TB | Prevents data skew or out-of-memory (OOM) errors in individual tasks. |
spark.sql.files.maxPartitionBytes |
128MB or 256MB |
Controls partition size during the read phase to avoid small file issues. |