Principal component analysis (PCA) is a multivariate statistical method used to explore the internal structures of multiple variables and how they correlate to each other based on a few principal components. You can use PCA to export a few principal components that are unrelated to each other from original variables. These principal components retain as much information about the original variables as possible and are used as new comprehensive metrics.
Limits
The Principal Component Analysis (PCA) component supports only data in the dense format. You can use the component for dimensionality and noise reduction.
Configure the component
You can use one of the following methods to configure the Principal Component Analysis (PCA) component.
Method 1: Configure the component on the pipeline page
| Tab | Parameter | Description |
| Fields Setting | Feature Columns | The columns that are selected from the input table for analysis. |
| Appended Columns | The columns that are appended to the table after dimensionality reduction. | |
| Parameters Setting | Data Size Ratio | The information retaining ratio after dimensionality reduction. |
| Feature Decomposition Mode | The method that is used to decompose features. Valid values:
| |
| Data Conversion Method | The method that is used to convert data types. Valid values:
| |
| Tuning | Lifecycle | The lifecycle of the output table. The value must be a positive integer. |
| Cores | The number of cores. This parameter is used with the Memory Size per Node (Unit: MB) parameter. The value must be a positive integer. Valid values: [1,9999]. | |
| Memory Size per Node (Unit: MB) | Unit: MB. The memory size of each core. The value must be a positive integer. Valid values: [1024,64 × 1024]. |
Method 2: Use PAI commands
PAI -name PrinCompAnalysis
-project algo_public
-DinputTableName=bank_data
-DeigOutputTableName=pai_temp_2032_17900_2
-DprincompOutputTableName=pai_temp_2032_17900_1
-DselectedColNames=pdays,previous,emp_var_rate,cons_price_idx,cons_conf_idx,euribor3m,nr_employed
-DtransType=Simple
-DcalcuType=CORR
-DcontriRate=0.9;| Parameter | Required | Description | Default value |
| inputTableName | Yes | The input table that is used for training. | No default value |
| selectedColNames | Yes | The columns that are selected from the input table for analysis. Separate multiple columns with commas (,). The columns of the INT or DOUBLE data type are supported. | No default value |
| eigOutputTableName | Yes | The output table that contains feature vectors and feature values. | No default value |
| princompOutputTableName | Yes | The output table after dimensionality and noise reduction of principal components. | No default value |
| transType | No | The method that is used to transform the original table to a PCA table. Valid values:
| Simple |
| calcuType | No | The method that is used to decompose features of the original table. Valid values:
| CORR |
| contriRate | No | The information retaining ratio after dimensionality reduction. Valid values: (0,1). | 0.9 |
| remainColumns | No | The fields that are retained from the original table after dimensionality reduction. | No default value |
| coreNum | No | The number of cores. This parameter is used with the memSizePerCore parameter. The value must be a positive integer. Valid values: [1,9999]. | Determined by the system |
| memSizePerCore | No | The memory size of each core. Unit: MB. The memory size of each core. The value must be a positive integer. Valid values: [1024,64 × 1024]. | Determined by the system |
| lifecycle | No | The lifecycle of the output table. The value must be a positive integer. | No default value |
Example
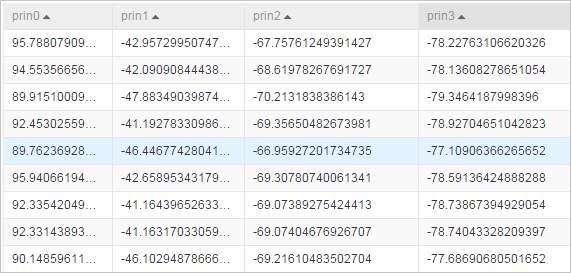
- Data table after dimensionality reduction
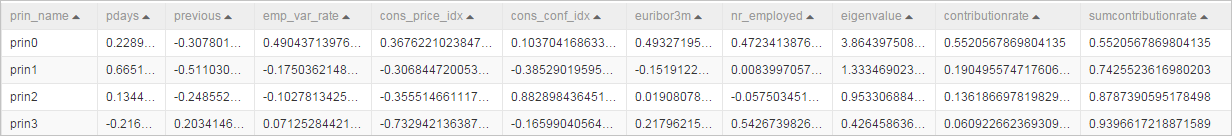
- Table that contains feature values and feature vectors