Use the Label Propagation Algorithm (LPA) to detect communities in social networks, knowledge graphs, and recommendation systems by iteratively assigning each vertex the most frequent label among its neighbors.
How it works
-
Graph clustering divides graphs into subgraphs based on topology, maximizing connections within subgraphs and minimizing connections between subgraphs.
-
The algorithm initializes each vertex with a unique label, then iteratively assigns each vertex the most frequent label among its neighbors. Iteration stops when all vertices have the most frequent label from their neighbors.
Configure the component
Method 1: Configure on pipeline page
Add the Label Propagation Clustering component to the pipeline canvas and configure the following parameters.
|
Tab |
Parameter |
Description |
|
Fields Setting |
Vertex Table: Vertex Column |
Vertex column name in the vertex table. |
|
Vertex Table: Weight Column |
Weight column name in the vertex table. |
|
|
Edge Table: Source Vertex Column |
Source vertex column name in the edge table. |
|
|
Edge Table: Target Vertex Column |
Target vertex column name in the edge table. |
|
|
Edge Table: Weight Column |
Weight column name in the edge table. |
|
|
Parameters Setting |
Maximum Iterations |
Maximum number of iterations. Default: 30. |
|
Tuning |
Workers |
Number of workers for parallel execution. Higher values increase parallelism but also communication costs. |
|
Memory Size per Worker (MB) |
Maximum memory per worker in MB. Default: 4096. If memory usage exceeds this value, an |
Method 2: Use PAI commands
Configure the component using PAI commands within the SQL Script component. For more information, see Scenario 4: Execute PAI commands within the SQL script component in the "SQL Script" topic.
PAI -name LabelPropagationClustering
-project algo_public
-DinputEdgeTableName=LabelPropagationClustering_func_test_edge
-DfromVertexCol=flow_out_id
-DtoVertexCol=flow_in_id
-DinputVertexTableName=LabelPropagationClustering_func_test_node
-DvertexCol=node
-DoutputTableName=LabelPropagationClustering_func_test_result
-DhasEdgeWeight=true
-DedgeWeightCol=edge_weight
-DhasVertexWeight=true
-DvertexWeightCol=node_weight
-DrandSelect=true
-DmaxIter=100;|
Parameter |
Required |
Default value |
Description |
|
inputEdgeTableName |
Yes |
No default value |
Input edge table name. |
|
inputEdgeTablePartitions |
No |
Full table |
Partitions to read from the input edge table. |
|
fromVertexCol |
Yes |
No default value |
Source vertex column name in the edge table. |
|
toVertexCol |
Yes |
No default value |
Target vertex column name in the edge table. |
|
inputVertexTableName |
Yes |
No default value |
Input vertex table name. |
|
inputVertexTablePartitions |
No |
Full table |
Partitions to read from the input vertex table. |
|
vertexCol |
Yes |
No default value |
Vertex column name in the vertex table. |
|
outputTableName |
Yes |
No default value |
Output table name. |
|
outputTablePartitions |
No |
No default value |
Partitions to write in the output table. |
|
lifecycle |
No |
No default value |
Lifecycle of the output table in days. |
|
workerNum |
No |
No default value |
Number of workers for parallel execution. Higher values increase parallelism but also communication costs. |
|
workerMem |
No |
4096 |
Maximum memory per worker in MB. Default: 4096. If memory usage exceeds this value, an |
|
splitSize |
No |
64 |
Data split size in MB. |
|
hasEdgeWeight |
No |
false |
Whether edges have weights. |
|
edgeWeightCol |
No |
No default value |
Weight column name in the edge table. |
|
hasVertexWeight |
No |
false |
Whether vertices have weights. |
|
vertexWeightCol |
No |
No default value |
Weight column name in the vertex table. |
|
randSelect |
No |
false |
Whether to randomly select when multiple labels have the same maximum frequency. |
|
maxIter |
No |
30 |
Maximum number of iterations. |
Example
-
Add the SQL Script component to the canvas and run the following SQL statements to generate training data.
drop table if exists LabelPropagationClustering_func_test_edge; create table LabelPropagationClustering_func_test_edge as select * from ( select '1' as flow_out_id,'2' as flow_in_id,0.7 as edge_weight union all select '1' as flow_out_id,'3' as flow_in_id,0.7 as edge_weight union all select '1' as flow_out_id,'4' as flow_in_id,0.6 as edge_weight union all select '2' as flow_out_id,'3' as flow_in_id,0.7 as edge_weight union all select '2' as flow_out_id,'4' as flow_in_id,0.6 as edge_weight union all select '3' as flow_out_id,'4' as flow_in_id,0.6 as edge_weight union all select '4' as flow_out_id,'6' as flow_in_id,0.3 as edge_weight union all select '5' as flow_out_id,'6' as flow_in_id,0.6 as edge_weight union all select '5' as flow_out_id,'7' as flow_in_id,0.7 as edge_weight union all select '5' as flow_out_id,'8' as flow_in_id,0.7 as edge_weight union all select '6' as flow_out_id,'7' as flow_in_id,0.6 as edge_weight union all select '6' as flow_out_id,'8' as flow_in_id,0.6 as edge_weight union all select '7' as flow_out_id,'8' as flow_in_id,0.7 as edge_weight )tmp ; drop table if exists LabelPropagationClustering_func_test_node; create table LabelPropagationClustering_func_test_node as select * from ( select '1' as node,0.7 as node_weight union all select '2' as node,0.7 as node_weight union all select '3' as node,0.7 as node_weight union all select '4' as node,0.5 as node_weight union all select '5' as node,0.7 as node_weight union all select '6' as node,0.5 as node_weight union all select '7' as node,0.7 as node_weight union all select '8' as node,0.7 as node_weight )tmp;Data structure
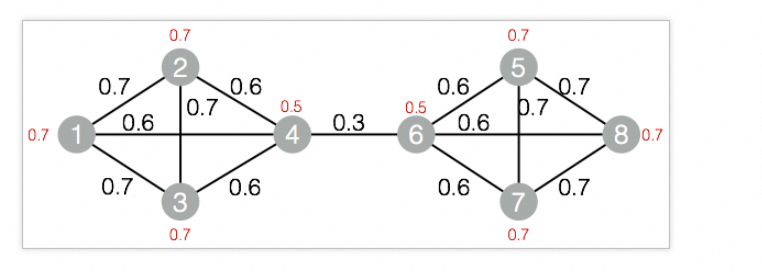
-
Add the SQL Script component to the canvas and run the following PAI commands.
drop table if exists ${o1}; PAI -name LabelPropagationClustering -project algo_public -DinputEdgeTableName=LabelPropagationClustering_func_test_edge -DfromVertexCol=flow_out_id -DtoVertexCol=flow_in_id -DinputVertexTableName=LabelPropagationClustering_func_test_node -DvertexCol=node -DoutputTableName=${o1} -DhasEdgeWeight=true -DedgeWeightCol=edge_weight -DhasVertexWeight=true -DvertexWeightCol=node_weight -DrandSelect=true -DmaxIter=100; -
Right-click the SQL Script component and choose View Data > SQL Script Output to view results.
| node | group_id | | ---- | -------- | | 1 | 3 | | 3 | 3 | | 5 | 7 | | 7 | 7 | | 2 | 3 | | 4 | 3 | | 6 | 7 | | 8 | 7 |