This topic uses Schrodinger as an example to describe how to calculate molecular structure by using an Elastic High Performance Computing (E-HPC) cluster.
Background information
Schrodinger provides a physics-based computational platform that integrates differentiated solutions for predictive modeling and data analysis to facilitate the exploration of the chemical space. The platform can be used in drug discovery and materials science in various fields, such as aerospace, energy, semiconductors, and electronic displays. For more information, see the official website of Schrodinger.
Preparations
Create an E-HPC cluster. For more information, see Create a cluster by using the wizard.
Configure the following parameters.
Parameter
Description
Hardware settings
Deploy a standard cluster, which consists of two management nodes, one compute node, and one logon node. Select GPU-accelerated instance types for compute nods. In this example, ecs.gn5-c4g1.xlarge is used.
Software settings
Deploy a CentOS 7.6 public image and the Slurm scheduler. Turn on VNC.
Create a cluster user. For more information, see Create a user.
The user is used to log on to the cluster, compile LAMMPS, and submit jobs. In this example, the following information is used to create the user:
Username: testuser
User group: sudo permission group
Install the VMD software. For more information, see Install software:
Download and install the Schrodinger software.
Schrodinger is paid software. For more information about how to purchase, download, install, and use Schrodinger software, see the official website of Schrodinger.
Download and install the CUDA driver on the compute nodes.
For more information about how to download and install the CUDA driver, see NVIDIA CUDA.
Step 1: Configure Schrodinger
Before you start to use Schrodinger for the first time, you need to configure the environment.
Log on to the cluster as the root user. For more information, see Log on to an E-HPC cluster.
Modify the configuration file of the Slurm scheduler.
vim /opt/schrodinger/queues/SLURM2.1/configIn the configuration file, specify REMOTE_LICENSE_SERVER=${ehpcname} to the server that you set when you register the license for the Slurm scheduler. Example:
QPATH=/usr/bin QPROFILE= QSUB=sbatch QDEL=scancel QSTAT=squeue LICENSE_CHECKING=yes REMOTE_LICENSE_SERVER=schrodinge_ehpc-testConfigure the hosts file of Schrodinger.
vim /opt/schrodinger/schrodinger.hostsSample content:
name: comp-gpu queue: SLURM2.1 qargs: --partition=comp --nodes=1 --ntasks=%NPROC% --mem=3G --gres=gpu:%NPROC% host: manager processors: 4 gpgpu: 0, NVIDIA P100
Step 2: Submit a job
Log on to the cluster as the cluster user. In this example, testuser is used. For more information, see Log on to an E-HPC cluster.
Run the following commands to download and decompress the sample file:
wget https://public-ehpc-package.oss-cn-hangzhou.aliyuncs.com/desmond_md_job_10-GPU.tar.gz tar xf desmond_md_job_10-GPU.tar.gzRun the following commands to submit the job:
cd desmond_md_job_10 ./desmond_md_job_10.shThe following example response indicates that the generated job ID is manager-0-646496c8.
JobId: manager-0-646496c8
Step 3: View the job result
Run the following command to view the job execution status:
squeueThe job information is displayed in the response. An
Rin theSTcolumn indicates that the job is running.JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON) 3 comp desmond_ testuser R 1:27 1 compute000NoteThe job takes about 10 minutes to run. After the job is run, you can run the
sacctcommand to check whether the job is completed.The job result is exported to the desmond_md_job_10-out.cms.
Use VNC to view the result of the job.
Enable VNC.
In the left-side navigation pane of the E-HPC console, click Cluster.
On the Cluster page, select a cluster. Choose More > VNC.
Use VNC to remotely connect to a visualization service. For more information, see Use VNC to manage a visualization service.
In the Virtualization Service dialog box of the cloud desktop, choose Application > System Tools > Terminal.
Run the
/opt/vmd/1.9.3/vmdto open Visual Molecular Dynamics (VMD).On the VMD Main page, choose File > New Molecule....
Click Browse... at the Filename to select the result file named desmond_md_job_10-out.cms.
In this example, testuser is used as the user. The path of the job result file is
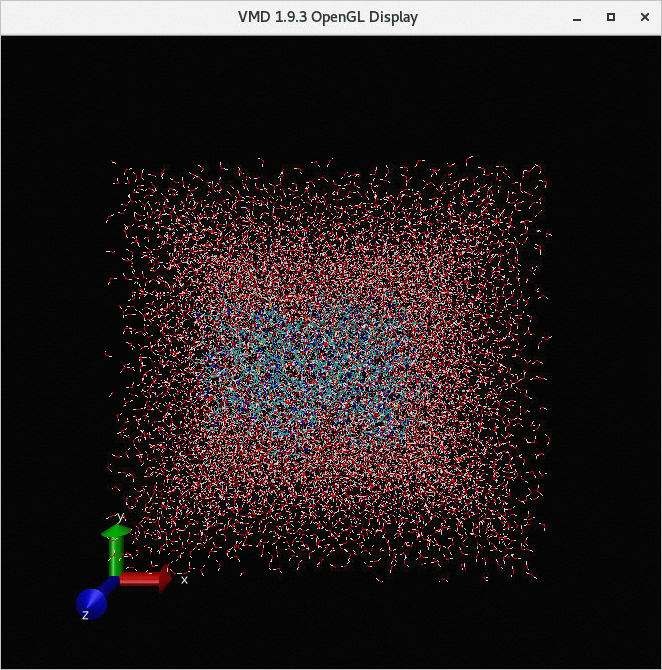
/home/testuser/desmond_md_job_10/desmond_md_job_10-out.cms.Click Load. You can view the result of the job in the VMD 1.9.3 OpenGL Display window.