PASE (PostgreSQL ANN search extension) is a high-performance vector similarity search plug-in for PolarDB for PostgreSQL. It implements two approximate nearest neighbor (ANN) algorithms — IVFFlat and Hierarchical Navigable Small World (HNSW) — so you can query high-dimensional vectors at speed directly from your database.
PASE does not extract or output feature vectors. Retrieve the feature vectors for your target entities first, then use PASE to run similarity searches across large vector datasets.
Prerequisites
Before you begin, ensure that you have:
A PolarDB for PostgreSQL cluster
A privileged account (required to run the SQL statements in this topic)
Basic familiarity with machine learning concepts and vector search
Usage notes
Index bloat: Run
select pg_relation_size('index_name');and compare the result with the table size. If the index is larger than the table and queries have slowed down, rebuild the index.Index drift after updates: Frequent data updates can reduce index accuracy. Rebuild the index regularly if you require 100% recall.
IVFFlat internal clustering: To create an IVFFlat index with internal centroids, set
clustering_type = 1and insert data into the table before creating the index.Multi-node elastic parallel queries: Only sequential search is supported for high-dimensional vectors in multi-node elastic parallel queries.
Choose an algorithm
PASE supports two ANN algorithms. The right choice depends on your dataset size, latency target, and recall requirements.
IVFFlat | HNSW | |
Best for | High-precision use cases (e.g., image comparison) | Large datasets with low-latency requirements |
Query latency target | Up to 100 ms | Up to 10 ms |
Dataset size | Any size | Tens of millions of vectors or more |
100% recall | Yes, when the query vector is in the candidate dataset | No; precision plateaus and cannot be increased further by tuning |
Build time | Fast | Slower |
Storage overhead | Low | Higher (proximity graph neighbors stored) |
Precision control | Fully controllable via parameters | Limited after a threshold |
IVFFlat
IVFFlat is a simplified version of the IVFADC algorithm. It clusters vectors using k-means, then searches only the clusters nearest to the query vector — skipping distant clusters to speed up the search. Precision scales with the number of clusters searched, so you can tune it directly.
How IVFFlat works:
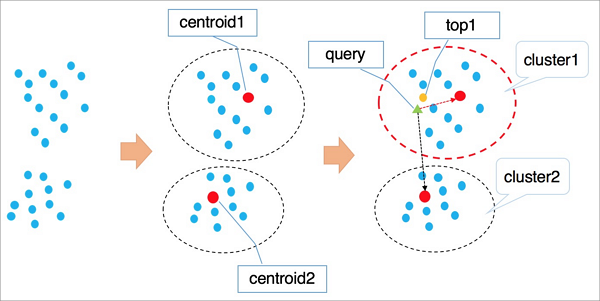
IVFFlat applies k-means clustering to group vectors into clusters, each with a centroid.
It identifies the
ncentroids nearest to the query vector.It traverses and ranks all vectors in those
nclusters and returns the nearestkvectors.
A larger value of n increases precision but also increases compute. Unlike IVFADC, which uses product quantization in phase 2 to reduce traversal cost (at the expense of precision), IVFFlat uses brute-force search — giving you direct control over the accuracy/performance tradeoff.
HNSW
HNSW builds a hierarchical multi-layer graph using the Navigable Small World (NSW) algorithm. Each layer acts as a panoramic skip list over the layer below, enabling fast traversal across large datasets.
How HNSW works:
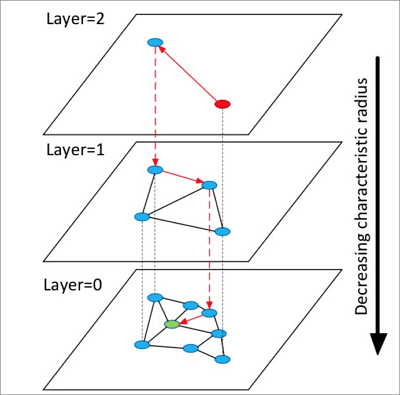
HNSW builds a structure with multiple layers (graphs). Each layer is a panorama and skip list of its lower layer.
Starting from a random element in the top layer, HNSW identifies neighbors and adds them to a fixed-length dynamic list ranked by distance.
It continues expanding neighbors, re-sorting the list after each addition and retaining only the top
k. Once the list stabilizes, HNSW descends to the next layer using the top element as the starting point.It repeats until it completes a search in the bottom layer.
After precision reaches a certain level, you cannot increase it further by reconfiguring parameters. HNSW also requires additional storage to persist proximity graph neighbors.
Enable PASE
Run the following statement to install the PASE extension:
CREATE EXTENSION pase;Calculate vector similarity
Before creating an index, you can calculate vector similarity directly using the <?> operator. PASE supports two construction methods.
The left vector must use thefloat4[]type and the right vector must use thePASEtype. Both vectors must have the same number of dimensions, or the operation returns a similarity calculation error.
PASE-type construction
The <?> operator computes the similarity between vectors. The PASE data type accepts up to three parameters:
Parameter 1: The right-side vector (
float4[])Parameter 2: Reserved; set to
0Parameter 3: Similarity method —
0for Euclidean distance,1for dot product (inner product)
SELECT ARRAY[2, 1, 1]::float4[] <?> pase(ARRAY[3, 1, 1]::float4[]) AS distance;
SELECT ARRAY[2, 1, 1]::float4[] <?> pase(ARRAY[3, 1, 1]::float4[], 0) AS distance;
SELECT ARRAY[2, 1, 1]::float4[] <?> pase(ARRAY[3, 1, 1]::float4[], 0, 1) AS distance;String-based construction
String-based construction uses colons (:) to separate parameters instead of function arguments:
SELECT ARRAY[2, 1, 1]::float4[] <?> '3,1,1'::pase AS distance;
SELECT ARRAY[2, 1, 1]::float4[] <?> '3,1,1:0'::pase AS distance;
SELECT ARRAY[2, 1, 1]::float4[] <?> '3,1,1:0:1'::pase AS distance;In '3,1,1:0:1', the first segment is the vector, the second is the reserved parameter (0), and the third is the similarity method (0 = Euclidean, 1 = dot product).
Normalization requirement
Euclidean distance: No normalization needed.
Dot product or cosine: Normalize the original vector first. For a normalized vector, the dot product equals the cosine value. For example, if the original vector is
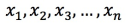 , it must satisfy:
, it must satisfy: 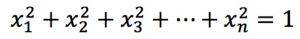 .
.
Create an index
IVFFlat index
CREATE INDEX ivfflat_idx ON vectors_table
USING
pase_ivfflat(vector)
WITH
(clustering_type = 1, distance_type = 0, dimension = 256, base64_encoded = 0, clustering_params = "10,100");Parameters
Parameter | Required | Default | Description |
| Yes | — | Clustering mode. |
| No |
| Similarity method. |
| Yes | — | Number of dimensions. Maximum: 512. |
| No |
| Vector encoding. |
| Yes | — | For external clustering: the directory of the centroid file. For internal clustering: |
`clustering_params` for internal clustering (`clustering_type = 1`)
Format: clustering_sample_ratio,k
clustering_sample_ratio: Sampling fraction with 1,000 as the denominator. Range: (0, 1000]. For example,1means a 1/1000 sampling ratio. A higher value increases accuracy but slows index creation. Keep total sampled records under 100,000.k: Number of centroids. A higher value increases accuracy but slows index creation. The recommended range is [100, 1000].
HNSW index
CREATE INDEX hnsw_idx ON vectors_table
USING
pase_hnsw(vector)
WITH
(dim = 256, base_nb_num = 16, ef_build = 40, ef_search = 200, base64_encoded = 0);Parameters
Parameter | Required | Default | Description |
| Yes | — | Number of dimensions. Maximum: 512. |
| Yes | — | Number of neighbors per element. A higher value increases accuracy but slows index creation and uses more storage. Recommended range: [16, 128]. |
| Yes | — | Heap length during index creation. A longer heap increases accuracy but slows creation. Recommended range: [40, 400]. |
| Yes |
| Heap length during queries. A larger value increases accuracy but reduces query performance. Can be overridden at query time. Start at |
| No |
| Vector encoding. |
Query vectors
Query with an IVFFlat index
The <#> operator is used for IVFFlat index queries. An ORDER BY clause is required for the index to take effect. Results are sorted in ascending order by distance.
SELECT id, vector <#> '1,1,1'::pase AS distance
FROM vectors_ivfflat
ORDER BY
vector <#> '1,1,1:10:0'::pase
ASC LIMIT 10;The query string '1,1,1:10:0' contains three parameters separated by colons:
Position | Value | Description |
1 |
| Query vector |
2 |
| Query accuracy — range: (0, 1000]. Higher = more accurate, lower query performance. |
3 |
| Similarity method: |
Query with an HNSW index
The <?> operator is used for HNSW index queries. An ORDER BY clause is required for the index to take effect. Results are sorted in ascending order by distance.
SELECT id, vector <?> '1,1,1'::pase AS distance
FROM vectors_ivfflat
ORDER BY
vector <?> '1,1,1:100:0'::pase
ASC LIMIT 10;The query string '1,1,1:100:0' contains three parameters separated by colons:
Position | Value | Description |
1 |
| Query vector |
2 |
| Query accuracy — range: (0, ∞). Higher = more accurate, lower query performance. Start at |
3 |
| Similarity method: |
Appendix
Calculate dot product (inner product)
Because the dot product of a normalized vector equals its cosine value, the following function works for both dot product and cosine similarity searches. It uses an HNSW index.
CREATE OR REPLACE FUNCTION inner_product_search(query_vector text, ef integer, k integer, table_name text) RETURNS TABLE (id integer, uid text, distance float4) AS $$
BEGIN
RETURN QUERY EXECUTE format('
select a.id, a.vector <?> pase(ARRAY[%s], %s, 1) AS distance from
(SELECT id, vector FROM %s ORDER BY vector <?> pase(ARRAY[%s], %s, 0) ASC LIMIT %s) a
ORDER BY distance DESC;', query_vector, ef, table_name, query_vector, ef, k);
END
$$
LANGUAGE plpgsql;Create an IVFFlat index from an external centroid file
This is an advanced feature. Upload an external centroid file to the specified server directory and reference it in clustering_params. The file format is:
Number of dimensions|Number of centroids|Centroid vector datasetExample:
3|2|1,1,1,2,2,2References
Hervé Jégou, Matthijs Douze, Cordelia Schmid. Product quantization for nearest neighbor search. IEEE.
Yu.A.Malkov, D.A.Yashunin. Efficient and robust approximate nearest neighbor search using Hierarchical Navigable Small World graphs. IEEE.