TairVector is an in-memory vector engine that supports real-time index updates and millisecond-level k-nearest neighbor (KNN) search. This tutorial shows how to build a molecular structure similarity search system on TairVector using Python and RDKit—a workflow common in AI-powered drug discovery and compound screening.
Background
Screening large compound libraries for structurally similar molecules is a core task in drug discovery. The typical approach converts each molecular structure from its SMILES representation into a fixed-length vector fingerprint, stores the fingerprints in a vector index, and then retrieves the k most similar structures for a query molecule.
TairVector stores all data in memory and supports real-time index updates, which gives it lower read and write latency than disk-based alternatives. Key characteristics for this use case:
| Characteristic | Description |
|---|---|
| In-memory storage | Keeps the entire molecular fingerprint index in RAM for low-latency access |
| Real-time index updates | New compounds can be indexed without rebuilding from scratch |
KNN search via TVS.KNNSEARCH |
Retrieves the top k most similar structures in milliseconds |
| Configurable k | Adjust the number of results returned to match your screening pipeline |
Prerequisites
Before you begin, make sure you have:
-
A Tair instance. Record its endpoint and password.
-
Python 3.8 or later
-
The following Python dependencies installed:
pip install numpy rdkit tair matplotlib
How it works
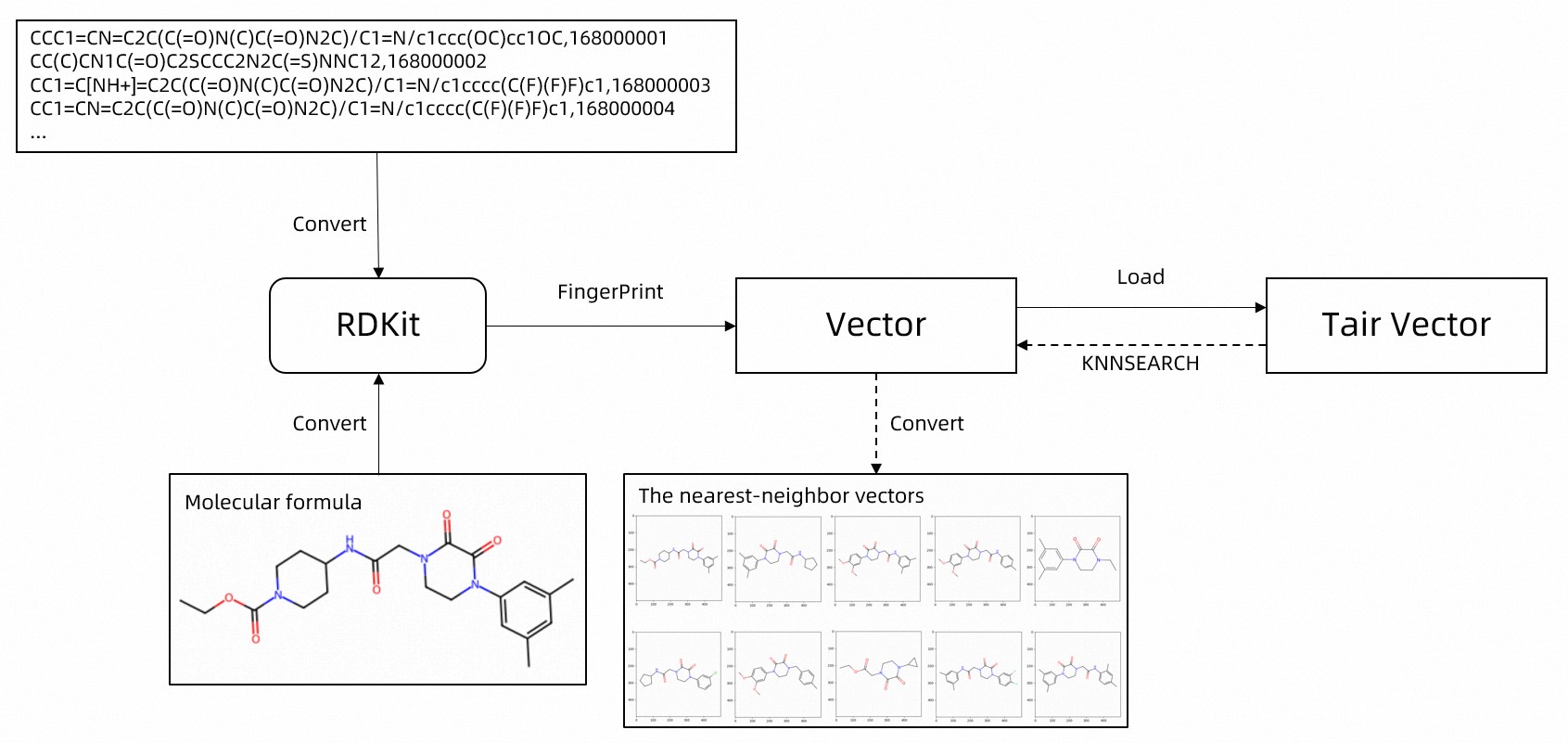
The end-to-end pipeline consists of five steps:
-
Download the molecular structure dataset in SMILES format.molecular structure dataset
-
Connect to a Tair instance.
-
Create a vector index to store the molecular fingerprints.
-
Convert each SMILES string to a 512-dimensional Morgan fingerprint vector and write it to the index.
-
Query the index for the k most similar structures to a target molecule.
All five steps are implemented in the sample code below. Each section explains the corresponding function.
Step 1: Prepare the dataset
Download the sample dataset from PubChem. It contains 11,012 compounds in Simplified Molecular Input Line Entry System (SMILES) format, with two columns: chemical formula and unique ID.
CCC1=CN=C2C(C(=O)N(C)C(=O)N2C)/C1=N/c1ccc(OC)cc1OC,168000001
CC(C)CN1C(=O)C2SCCC2N2C(=S)NNC12,168000002
CC1=C[NH+]=C2C(C(=O)N(C)C(=O)N2C)/C1=N/c1cccc(C(F)(F)F)c1,168000003
CC1=CN=C2C(C(=O)N(C)C(=O)N2C)/C1=N/c1cccc(C(F)(F)F)c1,168000004In a production environment, load a larger dataset to test TairVector's millisecond-level retrieval performance at scale.
If your data is in Structure-Data File (SDF) format from the PubChem FTP server, convert it to SMILES first using RDKit:
import sys
from rdkit import Chem
def converter(file_name):
mols = [mol for mol in Chem.SDMolSupplier(file_name)]
outname = file_name.split(".sdf")[0] + ".smi"
out_file = open(outname, "w")
for mol in mols:
smi = Chem.MolToSmiles(mol)
name = mol.GetProp("_Name")
out_file.write("{},{}\n".format(smi, name))
out_file.close()
if __name__ == "__main__":
converter(sys.argv[1])Step 2: Connect to a Tair instance
The get_tair() function establishes a connection to your Tair instance. Replace the placeholder values with your actual endpoint and password.
Avoid hardcoding credentials in your code. Store the endpoint and password in environment variables and read them at runtime using os.environ.get().from tair import Tair
def get_tair() -> Tair:
"""
Connect to the Tair instance.
host: The endpoint of the Tair instance.
port: The port number. Default is 6379.
password: The password of the default account. To connect with a custom account, use 'username:password'.
"""
tair: Tair = Tair(
host="r-bp1mlxv3xzv6kf****pd.redis.rds.aliyuncs.com",
port=6379,
db=0,
password="Da******3",
)
return tairStep 3: Create a vector index
The create_index() function creates a vector index named MOLSEARCH_TEST. If the index already exists, it skips creation.
def create_index():
"""
Create a vector index named MOLSEARCH_TEST with the following parameters:
- Dimensions: 512 (matches the Morgan fingerprint bit length)
- Distance metric: L2 (Euclidean distance)
- Index algorithm: HNSW (Hierarchical Navigable Small World)
"""
ret = tair.tvs_get_index(INDEX_NAME)
if ret is None:
tair.tvs_create_index(INDEX_NAME, 512, distance_type=DistanceMetric.L2, index_type="HNSW")
print("create index done")The vector dimension is set to 512 to match the output ofsmiles_to_vector(), which usesAllChem.GetMorganFingerprintAsBitVect(mol, radius=2, nBits=512*8). The L2 distance metric measures Euclidean distance between fingerprint vectors; a lower score indicates greater structural similarity.
Step 4: Write molecular structure data
The do_load() function reads the SMILES dataset, converts each entry to a 512-dimensional vector using the smiles_to_vector() function, and writes the result to TairVector with TVS.HSET via insert_data(). Writes are batched in groups of 10 and submitted concurrently using ThreadPoolExecutor.
Each entry is stored in the index as follows:
| Field | Value | Example |
|---|---|---|
| Key | Unique compound ID | 168000001 |
| Vector | 512-dimensional Morgan fingerprint | — |
Attribute: smiles |
Original chemical formula string | CCC1=CN=C2... |
from concurrent.futures import ThreadPoolExecutor
from rdkit.Chem import AllChem
from rdkit import DataStructs, Chem
from rdkit import RDLogger
RDLogger.DisableLog('rdApp.*')
def do_load(file_path):
num = 0
lines = []
with open(file_path, 'r') as f:
for line in f:
if line.find("smiles") >= 0:
continue
lines.append(line)
if len(lines) >= 10:
parallel_submit_lines(lines)
num += len(lines)
lines.clear()
if num % 10000 == 0:
print("load num", num)
if len(lines) > 0:
parallel_submit_lines(lines)
print("load done")
def parallel_submit_lines(lines):
with ThreadPoolExecutor(len(lines)) as t:
for line in lines:
t.submit(handle_line, line=line)
def handle_line(line):
if line.find("smiles") >= 0:
return
parts = line.strip().split(',')
try:
ids = parts[1]
smiles = parts[0]
vec = smiles_to_vector(smiles)
insert_data(ids, smiles, vec)
except Exception as result:
print(result)
def smiles_to_vector(smiles):
"""Convert a SMILES string to a 512-dimensional Morgan fingerprint vector."""
mols = Chem.MolFromSmiles(smiles)
fp = AllChem.GetMorganFingerprintAsBitVect(mols, 2, 512 * 8)
hex_fp = DataStructs.BitVectToFPSText(fp)
vec = list(bytearray.fromhex(hex_fp))
return vec
def insert_data(id, smiles, vector):
"""Write the vector and chemical formula to TairVector using TVS.HSET."""
attr = {'smiles': smiles}
tair.tvs_hset(INDEX_NAME, id, vector, **attr)Step 5: Search for similar molecular structures
The do_search() function takes a query molecule in SMILES format and an integer k, converts the query to a fingerprint vector, and runs TVS.KNNSEARCH against the MOLSEARCH_TEST index. It then fetches the chemical formula for each result using TVS.HMGET.
def do_search(search_smiles, k):
"""
Query the index for the k most similar molecular structures.
Returns unique IDs, L2 distance scores, and chemical formulas.
"""
vector = smiles_to_vector(search_smiles)
result = tair.tvs_knnsearch(INDEX_NAME, k, vector)
print("The 10 molecular structures most similar to the query target are as follows:")
for key, value in result:
similar_smiles = tair.tvs_hmget(INDEX_NAME, key, "smiles")
print(key, value, similar_smiles)Complete sample code
ReplaceD:\Test\Compound_168000001_168500000.smiin thedo_load()call with the actual path to your downloaded dataset file.
import os
import sys
from tair import Tair
from tair.tairvector import DistanceMetric
from rdkit.Chem import Draw, AllChem
from rdkit import DataStructs, Chem
from rdkit import RDLogger
from concurrent.futures import ThreadPoolExecutor
RDLogger.DisableLog('rdApp.*')
def get_tair() -> Tair:
"""
Connect to the Tair instance.
host: The endpoint of the Tair instance.
port: The port number. Default is 6379.
password: The password of the default account. To connect with a custom account, use 'username:password'.
"""
tair: Tair = Tair(
host="r-bp1mlxv3xzv6kf****pd.redis.rds.aliyuncs.com",
port=6379,
db=0,
password="Da******3",
)
return tair
def create_index():
"""
Create a vector index named MOLSEARCH_TEST:
- Dimensions: 512
- Distance metric: L2
- Index algorithm: HNSW
"""
ret = tair.tvs_get_index(INDEX_NAME)
if ret is None:
tair.tvs_create_index(INDEX_NAME, 512, distance_type=DistanceMetric.L2, index_type="HNSW")
print("create index done")
def do_load(file_path):
"""
Read the SMILES dataset, extract vector features, and write data to TairVector.
Data is stored as: key=compound ID, vector=512-dim fingerprint, attribute smiles=chemical formula.
"""
num = 0
lines = []
with open(file_path, 'r') as f:
for line in f:
if line.find("smiles") >= 0:
continue
lines.append(line)
if len(lines) >= 10:
parallel_submit_lines(lines)
num += len(lines)
lines.clear()
if num % 10000 == 0:
print("load num", num)
if len(lines) > 0:
parallel_submit_lines(lines)
print("load done")
def parallel_submit_lines(lines):
with ThreadPoolExecutor(len(lines)) as t:
for line in lines:
t.submit(handle_line, line=line)
def handle_line(line):
if line.find("smiles") >= 0:
return
parts = line.strip().split(',')
try:
ids = parts[1]
smiles = parts[0]
vec = smiles_to_vector(smiles)
insert_data(ids, smiles, vec)
except Exception as result:
print(result)
def smiles_to_vector(smiles):
"""Convert a SMILES string to a 512-dimensional Morgan fingerprint vector."""
mols = Chem.MolFromSmiles(smiles)
fp = AllChem.GetMorganFingerprintAsBitVect(mols, 2, 512 * 8)
hex_fp = DataStructs.BitVectToFPSText(fp)
vec = list(bytearray.fromhex(hex_fp))
return vec
def insert_data(id, smiles, vector):
"""Write the vector and chemical formula to TairVector using TVS.HSET."""
attr = {'smiles': smiles}
tair.tvs_hset(INDEX_NAME, id, vector, **attr)
def do_search(search_smiles, k):
"""
Query the index for the k most similar molecular structures.
Uses TVS.KNNSEARCH to find the nearest neighbors, then TVS.HMGET to retrieve their formulas.
"""
vector = smiles_to_vector(search_smiles)
result = tair.tvs_knnsearch(INDEX_NAME, k, vector)
print("The 10 molecular structures most similar to the query target are as follows:")
for key, value in result:
similar_smiles = tair.tvs_hmget(INDEX_NAME, key, "smiles")
print(key, value, similar_smiles)
if __name__ == "__main__":
# Connect to Tair and create the molecular structure index.
tair = get_tair()
INDEX_NAME = "MOLSEARCH_TEST"
create_index()
# Load the sample dataset.
do_load("D:\Test\Compound_168000001_168500000.smi")
# Query the 10 structures most similar to the target compound.
do_search("CCOC(=O)N1CCC(NC(=O)CN2CCN(c3cc(C)cc(C)c3)C(=O)C2=O)CC1", 10)Results
A successful run produces output similar to the following:
create index done
load num 10000
load done
The 10 molecular structures most similar to the query target are as follows:
b'168000009' 0.0 ['CCOC(=O)N1CCC(NC(=O)CN2CCN(c3cc(C)cc(C)c3)C(=O)C2=O)CC1']
b'168003114' 29534.0 ['Cc1cc(C)cc(N2CCN(CC(=O)NC3CCCC3)C(=O)C2=O)c1']
b'168000210' 60222.0 ['COc1ccc(N2CCN(CC(=O)Nc3cc(C)cc(C)c3)C(=O)C2=O)cc1OC']
b'168001000' 61123.0 ['COc1ccc(N2CCN(CC(=O)Nc3ccc(C)cc3)C(=O)C2=O)cc1OC']
b'168003038' 64524.0 ['CCN1CCN(c2cc(C)cc(C)c2)C(=O)C1=O']
b'168003095' 67591.0 ['O=C(CN1CCN(c2cccc(Cl)c2)C(=O)C1=O)NC1CCCC1']
b'168000396' 70376.0 ['COc1ccc(N2CCN(Cc3ccc(C)cc3)C(=O)C2=O)cc1OC']
b'168002227' 71121.0 ['CCOC(=O)CN1CCN(C2CC2)C(=O)C1=O']
b'168000441' 73197.0 ['Cc1cc(C)cc(NC(=O)CN2CCN(c3ccc(F)c(F)c3)C(=O)C2=O)c1']
b'168000561' 73269.0 ['Cc1cc(C)cc(N2CCN(CC(=O)Nc3ccc(C)cc3C)C(=O)C2=O)c1']Each row contains:
| Column | Description |
|---|---|
| Column 1 | Compound ID |
| Column 2 | L2 distance score (lower means more similar) |
| Column 3 | Chemical formula |
The result with score 0.0 is an exact match for the query molecule.
To visualize the retrieved structures as molecular images, use the following code:
import numpy
from rdkit.Chem import Draw
from rdkit import Chem
import matplotlib.pyplot as plt
def to_images(data):
imgs = []
for smiles in data:
mol = Chem.MolFromSmiles(smiles)
img = Chem.Draw.MolToImage(mol, size=(500, 500))
imgs.append(img)
plt.imshow(img)
plt.show()
return imgs
if __name__ == "__main__":
images = to_images(["CCOC(=O)N1CCC(NC(=O)CN2CCN(c3cc(C)cc(C)c3)C(=O)C2=O)CC1"])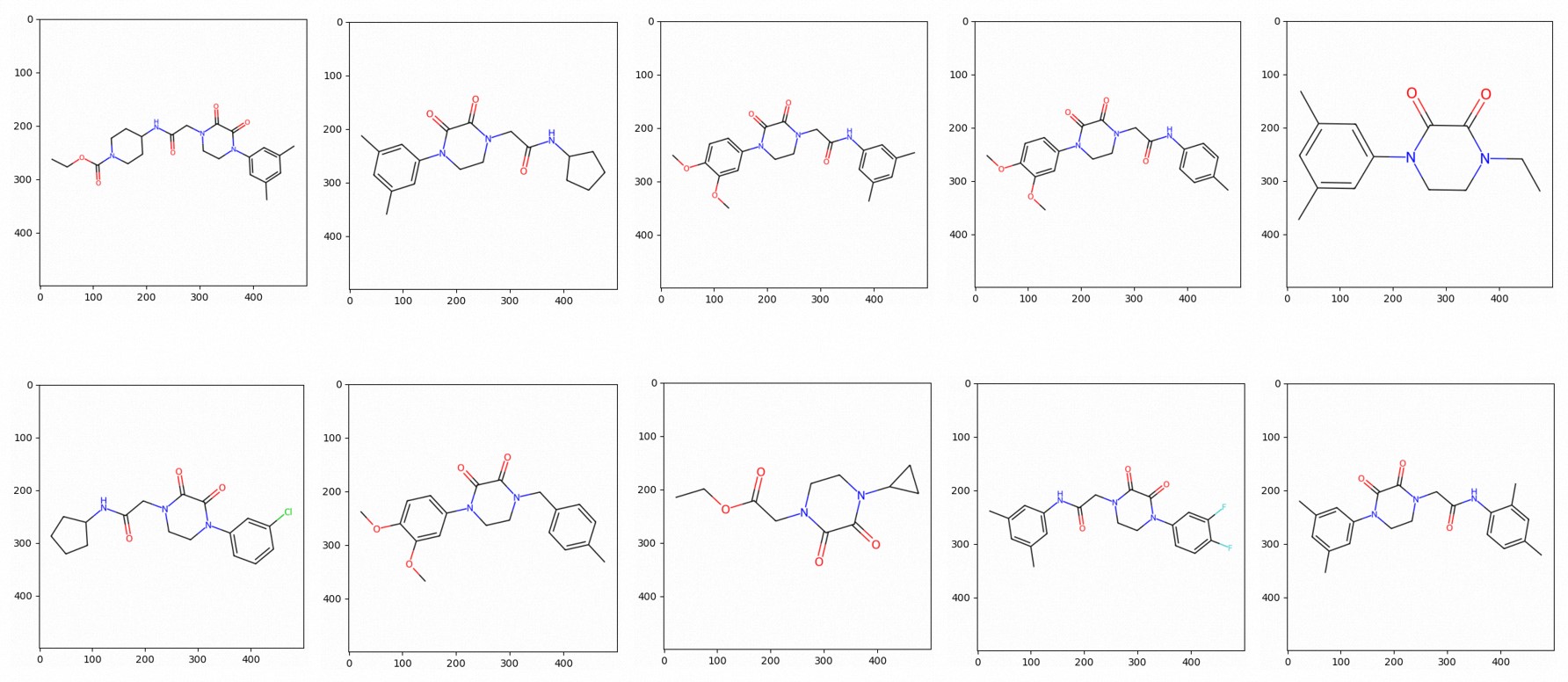
What's next
-
Load your full compound library and test retrieval performance at scale.
-
Adjust the
kparameter indo_search()to control how many candidates the screening pipeline returns.